Subscription packages, user molecule upload, docking visualization and more!
It has been a while since the last major mcule release, but now it is time for a big announcement! We are really proud of the outcome, so here are the new features:
1. Subscription packages
OK, here is something brand new in this field: you can now hire a single tool for a single project! This means that in contrast to the typical annual licensing schemes of modelling software companies we offer our packages for 1, 3, 6, and 12 month periods. You don't have the budget for maintaining an annual license for a tool? No problem, subscribe only when you really need it. Subscribing is super-easy: can be done in 1 minute!
We have signed partnership agreements with
BioSolveIT and
ChemAxon about distributing their products at
mcule.com. As a result you can now subscribe for drug discovery tools of these software companies
here. Besides, we are also introducing two mcule packages, but let's go step by step.
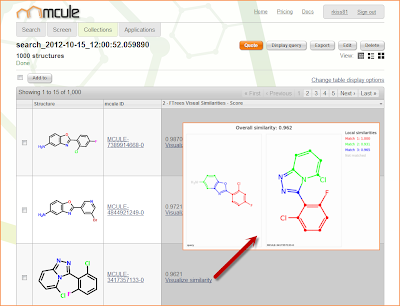
FTrees Visual Similarities is a great tool for scaffold hopping. It not only identifies diverse novel active scaffolds but it also tells why the query and target molecules are similar.
Physicochemical properties calculated by the popular Calculator plugins of ChemAxon are widely used and are known to give excellent correlation with experimental data. More than 100 precalculated
ChemAxon properties for the whole mcule database are available in the "Properties Access" package option (Academic users: it's free, so go to the
Pricing page, apply, and we will switch it on for you). To calculate ChemAxon properties with all possible settings, you should subscribe for the "Properties Access and Calculator" package option.
First mcule package is:
Database Access. It gives you access to two things: (i)
Product filter, and (ii) exporting chemical supplier data. The
Product filter is very useful to filter by supplier names, catalogs, stock availability, price, delivery time, etc. For example it can help to filter out unreliable suppliers and virtual compounds.
Second mcule package:
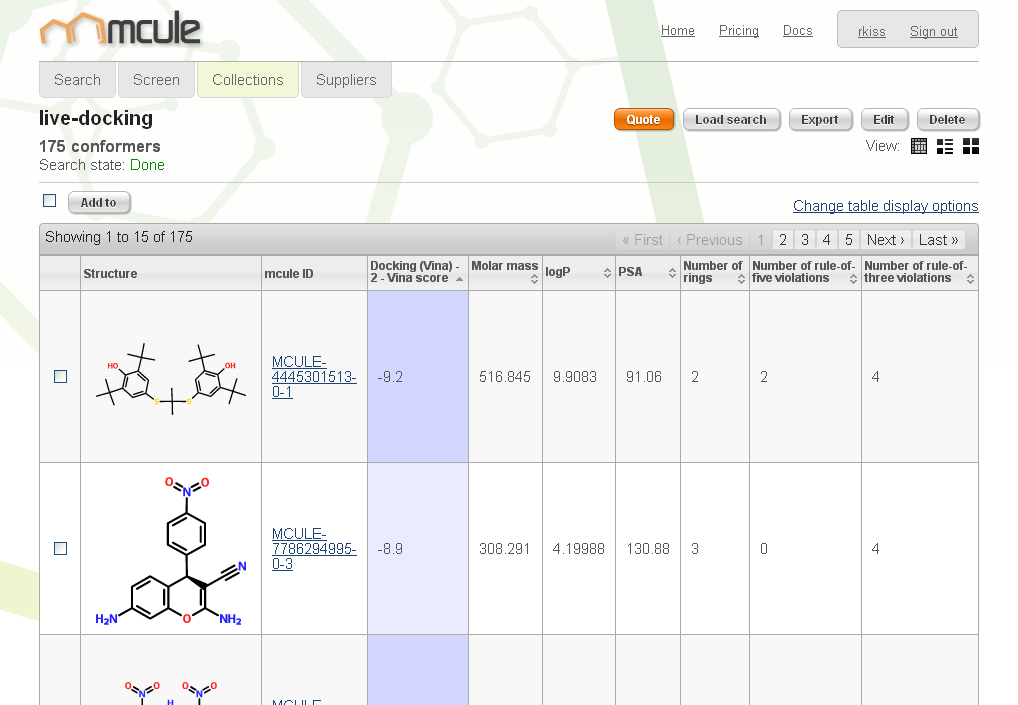
Docking (Vina). The
100k package option allows you to dock up to 100,000 molecules each month. We are planning to introduce a 1M package later on.
For more information about the subscription packages and package options check out our
Pricing page. To learn more about the individual drug discovery tools available at
mcule.com, check the
mcule documentation.
2. User molecule upload
Go to the "Collections" tab, and you will find a new button: "
Molecule upload". From now on, all the cool stuff available at
mcule.com are not limited for the mcule database. If you have your in-house database or have designed some virtual libraries you can upload them and run the searches and screens on your own libraries. To learn more about user collections, again, check the
mcule documentation.
3. Docking visualization
We have found and implemented a great Javascript 3D visualizer:
GLMol. After docking, there will appear a "
Visualize pose" link, which will display something like this:
Residue labeling has been also implemented. We continue working to further improve this visualization, but the user experience of the current version is already better than any WebGL molecule viewer (of course I'm biased, but still...). To get the best out of this, we recommend to run mcule in Chrome browser.
4. New tab: "Applications"
Under this tab, we will introduce a lot of useful stuff. First:
Property calculator. Draw a molecule or enter its identifier, then click on "Calculate" and you will see all mcule properties listed immediately. This works for molecules not in the mcule database too.
5. Documentation
I have already referred to our
documentation several times in this post. This is because it has been significantly extended and you can find most of the basic information about the mcule system there together with some nice screenshots. If you get in trouble you might also want to check the
FAQ page, where the most typical questions and answers are listed.
6. Flexibility
This is not as obvious as it should (or it is probably better if you don't notice anything of this), but we have completely redesigned the screening workflow technology behind the scenes. Why? To be able to get it to the next step: scale the system up to the sky. Now you can really put any workflow step after/before anything, and this makes the mcule technology really unique. We integrate this whole bunch of tools so flexibly, that will enable you setting whatever workflow you want.
What's next?
A lot of stuff as always, but I don't think docking a few million compounds will remain an issue too long. Only for the very few people around the world who are not mcule users yet. Do you know any? Please tell them to join
mcule! We honor the loyalty of our users especially those who bring new people. Soon you will find out how much!